Molecular Bonding Language
4 posters
Page 2 of 2
Page 2 of 2 •  1, 2
1, 2
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Ahh, DNA. Yes, I've had a go at building the DNA building blocks (GATC) before (many years ago) and they are complicated, for sure. It would be an interesting exercise in MBL. In the very least, it is a hell-of-a-lot easier than how I was doing it back then. You should start a new thread in the Chemistry section for it and we can at least work through the molecular structure of them.
With respect to a database, I have one and you are welcome to it, but I don't know if it will be exactly what you want. I designed a data model to store enough information for me to generate the models for AtomicViewer. I wasn't thinking about querying it for atomic information which I would probably design differently. However, it might be possible to use this database in order to generate a new one with higher level information. It really depends on your needs.
I had not thought about a database populated by MBL expressions before and it seems like an easy thing to do. What I mean is that I create an MBL Renderer that writes or executes the SQL to populate the database. Instead of a renderer that creates 3D models, this one generates SQL for a certain data model. I just need to know either what that data model is, or what it has to accomplish so that I can create it. The SQLRenderer is fairly easy to write unless the data model is too far removed from its concepts.
I might have a go at designing an almost direct representation of MBL as a database schema. It will pretty much be a database version of the Abstract Syntax Tree I've talked about above. Storing the raw data that an MBL expression declares. I'm not sure how useful that will be, but it might be a good start to other things. In the very least, it is easy to do.
With respect to a database, I have one and you are welcome to it, but I don't know if it will be exactly what you want. I designed a data model to store enough information for me to generate the models for AtomicViewer. I wasn't thinking about querying it for atomic information which I would probably design differently. However, it might be possible to use this database in order to generate a new one with higher level information. It really depends on your needs.
I had not thought about a database populated by MBL expressions before and it seems like an easy thing to do. What I mean is that I create an MBL Renderer that writes or executes the SQL to populate the database. Instead of a renderer that creates 3D models, this one generates SQL for a certain data model. I just need to know either what that data model is, or what it has to accomplish so that I can create it. The SQLRenderer is fairly easy to write unless the data model is too far removed from its concepts.
I might have a go at designing an almost direct representation of MBL as a database schema. It will pretty much be a database version of the Abstract Syntax Tree I've talked about above. Storing the raw data that an MBL expression declares. I'm not sure how useful that will be, but it might be a good start to other things. In the very least, it is easy to do.
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Cr6, what databases do you like to work in? I use MySQL at home, but often use Oracle, SQLServer and Postgres at work. I have also used SQLite and Apache Derby for small projects. The reason I ask is that writing DDL to populate a schema requires some way of saving row IDs so they can be used in child tables. Each database tends to do it differently. For MySQL I use session variables which are really nice to use. For example, I might do something like this:
The generated script is then run inside your favorite database app such as MySQL Workbench or Oracle SQLDeveloper or PGAdmin, etc.
- Code:
-- get the largest ID for a table and store it in a session variable
select (@chainId:=max(id)) from mbldb.chain;
select (@chainElementId:=max(id)) from mbldb.chain_element;
-- Create a chain
set @chainId := @chainId + 1;
insert into mbldb.chain (id, is_array, type) values (@chainId, 'N', NULL);
-- Create element for the chain.
set @chainElementId := @chainElementId + 1;
insert into mbldb.chain_element (id, chain_id, symbol, is_flipped, rotate, north, south, ordinal) values (@chainElementId, @chainId, 'O', 'N', 0, NULL, NULL, 1);
The generated script is then run inside your favorite database app such as MySQL Workbench or Oracle SQLDeveloper or PGAdmin, etc.
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Hey Nevyn, I actually work in SQL Server 2012+ mostly but do work in Oracle/MySQL to some extent. SQL Server 2016+ stores JSON natively so I would be looking at this for making a tabular version of Molecules. This was just a thought on doing ML on tables/tabular forms of the C.F. MBL if at all possible. I just have hunch that properties could be derived from it for automating structures. Might just be too much imagination at this point as well.
Last edited by Cr6 on Sun Jul 15, 2018 1:47 am; edited 1 time in total
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Hi Nevyn, I'm kinda stuck here (diving into MBL a bit) and thought posting here might be helpful for future references and newcomers too. My molecule so far:
ring w 45:(H)6{OH}

My question is, how do I get that Hydrogen on the end of the Oxygen to "turn" and face the O's charge field? I can't seem to get it to rotate on the proper axis. I tried:
ring w 45:(H)6{OH*90}
...and it rotates funny. What am I doing wrong?
Ultimately, I'm trying to generate THC obviously. Hope that doesn't deter anyone from the conversation or helping me out a little, though I understand if some people don't want to touch the topic.
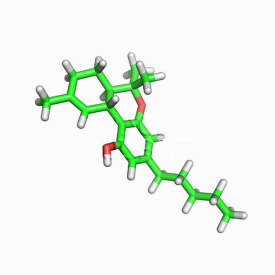
ring w 45:(H)6{OH}

My question is, how do I get that Hydrogen on the end of the Oxygen to "turn" and face the O's charge field? I can't seem to get it to rotate on the proper axis. I tried:
ring w 45:(H)6{OH*90}
...and it rotates funny. What am I doing wrong?
Ultimately, I'm trying to generate THC obviously. Hope that doesn't deter anyone from the conversation or helping me out a little, though I understand if some people don't want to touch the topic.

Jared Magneson- Posts : 525
Join date : 2016-10-11
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
It wouldn't turn, it should be closer and next to the proton in the Oxygen atom. It is a limitation of the MBL Renderer. It doesn't push the atoms together, it just sits them right next to each other.
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
You've picked a pretty difficult molecule to work with. There are bonds in that that I am not sure MBL can handle yet. Those connected rings are an issue, for sure. If you have trouble, just call out, because it might be MBL that is the problem.
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Yeah, I mean I knew it was a beast and perhaps I overreached here to start with. I'm gonna back up and work on a few much less complex molecules and learn the language better. It's really cool and I'm thankful you spent the time to take it even this far!
Jared Magneson- Posts : 525
Join date : 2016-10-11
 Re: Molecular Bonding Language
Re: Molecular Bonding Language
Thanks, Nevyn. It's obviously not critical, I was just trying to figure out from a mechanical perspective why this molecule is so unique. I guess it just has a lot of loose ends to bond with other substances? I mean, there's nothing "magical" about Oxygen, Hydrogen, and Carbon on its own. But it's been called among the most complex natural molecule occurring in plants, so I always wondered if that was true.
Just an aside, nothing to worry about here. It's fun to study and MBL really helps visualize things.
Just an aside, nothing to worry about here. It's fun to study and MBL really helps visualize things.
Jared Magneson- Posts : 525
Join date : 2016-10-11
Page 2 of 2 •  1, 2
1, 2
 Similar topics
Similar topics» Miles Periodic Table with Standard Periodic Table reference
» Atomic Modeling Language
» Spinning Particle Language
» Natural Language Processing (NLP) and newer algorithms
» Mass Defect May Help Explain Bonding & Gravity
» Atomic Modeling Language
» Spinning Particle Language
» Natural Language Processing (NLP) and newer algorithms
» Mass Defect May Help Explain Bonding & Gravity
Page 2 of 2
Permissions in this forum:
You cannot reply to topics in this forum|
|
|
